Introduction¶
Show how the sausage is made!
Wurst is a python package for linking and modifying industrial ecology models, with a focus on sparse matrices in life cycle assessment. Current development focuses on modifying the ecoinvent LCI database with scenario data from various data sources, using Brightway2 as the data backend.
See also the separate wurst examples repository.
A wurst model run will typically consist of the following steps:
- Load data from several sources
- Modify the LCI data
- Write the modified LCI data to some storage mechanism
Wurst supports the following generic modification types:
- Change the input material efficiency and associated emissions (
change_exchanges_by_constant_factor) - Change specific emissions separate from general efficiency improvements
- Change the relative shares of inputs (including adding new inputs) into markets
- Separate a global dataset into separate regions
In general, a modification function will include the following steps:
- Filter the LCI database by name, unit, location, etc. to get the subset of activities to modify
- Filter the external data source to get the relevant data used for modifications
- Change the values of some of the exchanges in the filtered LCI database using the filtered external data
Installation¶
Download and install miniconda, create and activate a new environment, and then install:
conda install -y -q -c conda-forge -c cmutel -c haasad -c konstantinstadler brightway2 jupyter wurst
Documents versus matrices¶
Inventory matrices can be modified by multiplying or adding vectors, as in the Themis methodology paper. Wurst takes a different approach - it treats each activity (column in the technosphere matrix) as a document with metadata and a list of exchanges which can be modified as desired. This approach allows for both flexibility (e.g. the number of rows and columns are not fixed) and simpler code (no need for an indirection layer to row and column indices). So, instead of constructing a vector and using it directly, wurst would prefer to write a function like:
import wurst as w
def scale_biosphere_exchanges_by_delta(ds, delta):
# Not directly related to fuel inputs
exclude_list = [
'Methane, fossil', 'Sulfur dioxide',
'Carbon monoxide, fossil',
'Nitrogen oxides', 'Dinitrogen monoxide', 'Particulates'
]
for exc in w.biosphere(ds, w.doesnt_contain_any('name', exclude_list)):
# Modifies in place
w.rescale_exchage(exc, delta)
Internal data format¶
The internal data format for Wurst is a subset of the implied internal format for Brightway2.
{
'database': str,
'code': str,
'name': str,
'reference product': str,
'location': str,
'unit': str,
'classifications': [tuple],
'categories': tuple or None,
'comment': str,
'parameters': {'parameter name (str)': float},
'exchanges': [
{
'amount': float,
'categories': list, # only for biosphere flows
'type': str, # biosphere, techosphere, production
'name': str,
'database': str,
'product': str,
'unit': str,
'location': str,
'input': tuple, # only if from external database
'uncertainty type': int, # optional
'loc': float, # optional
'scale': float, # optional
'shape': float, # optional
'minimum': float, # optional
'maximum': float, # optional
'production volume': float # optional
'pedigree': { # optional
'completeness': int,
'further technological correlation': int,
'geographical correlation': int,
'reliability': int,
'temporal correlation': int
},
}
]
}
An example classification:
('ISIC rev.4 ecoinvent', '1050:Manufacture of dairy products')
Searching and filtering¶
Wurst provides helper functions to make searching and filtering easier. These filter functions are designed to be used with get_many and get_one; here is an example:
nuclear_generation = get_many(
lci_database,
contains('name', 'nuclear'),
contains('name', 'electricity'),
equals('unit', 'kilowatt hour'),
exclude(contains('name', 'aluminium')),
exclude(contains('name', 'import'))
)
It is also OK to write a generator function that does the same thing:
nuclear_generation = (
ds for ds in lci_database
if 'nuclear' in ds['name']
and 'nuclear' in ds['name']
and ds['unit'] == 'kilowatt hour'
and 'aluminium' not in ds['name']
and 'import' not in ds['name']
)
The difference between the styles is ultimately a question of personal preference. For many people, list and generator expressions are more pythonic; in the specific case of wurst, using helper functions that are composable and reusable may allow you to not repeat yourself as often. There will also be times when the helper functions in wurst are not good enough for a specific search. In any case bear in mind the following general guidelines:
- Always manually check the results of your filtering functions before using them! The world is a complicated place, and our data sources reflect that complexity with unexpected or inconsistent elements.
- It is strongly recommended to use generator instead of list comprehensions, i.e.
(x for x in foo)instead of[x for x in foo].
For more information, see the introduction notebook, API documentation for searching, and: itertools, functools, toolz libraries.
Exchange iterators¶
The technosphere, biosphere, and production functions will return generators for exchanges with their respective exchange types.
Wurst also provides reference_product(dataset), which will return the single reference product for a dataset. If zero or multiple products are available, it will raise an error.
Transformations¶
-
wurst.transformations.activity.change_exchanges_by_constant_factor(ds, value, technosphere_filters=None, biosphere_filters=None)¶ Change some or all inputs and biosphere flows by a constant factor.
dsis a dataset document.valueis a number. Existing exchange amounts will be multiplied by this number.technosphere_filtersis an iterable of filter functions. Optional.biosphere_filtersis an iterable of filter functions. Optional.
Returns the altered dataset. The dataset is also modified in place, so the return value can be ignored.
Example: Changing coal dataset to reflect increased fuel efficiency
import wurst as w apct_products = w.either( w.equals('name', 'market for NOx retained'), w.equals('name', 'market for SOx retained'), ) generation_filters = [ w.either(w.contains('name', 'coal'), w.contains('name', 'lignite')), w.contains('name', 'electricity'), w.equals('unit', 'kilowatt hour'), w.doesnt_contain_any('name', [ 'market', 'aluminium industry', 'coal, carbon capture and storage' ]) ] fuel_independent = w.doesnt_contain_any('name', ( 'Methane, fossil', 'Sulfur dioxide', 'Carbon monoxide, fossil', 'Nitrogen oxides', 'Dinitrogen monoxide', 'Particulates' )) for ds in w.get_many(data, generation_filters): change_exchanges_by_constant_factor( ds, 0.8, # Or whatever from input data [w.exclude(apct_products)], [fuel_independent] )
Unlinking and Re-linking¶
Exchanges are considered “linked” if their input flows are already resolved to point to a certain producing activity. In Brightway2, this link is the field “input”, whose value takes the form ('database name', 'unique code'). Wurst uses the same convention - the input field is used to uniquely identify an activity that produces the exchange flow (biosphere flows are also considered activities).
The output field is not needed - this is the activity in question, which consumes the input flow. Production exchanges will have the same value in input and output.
The default Brightway2 importer will remove the input field for exchanges which are provided by another activity in the same set of input datasets. Instead of an input field, the exchange will have an activity name, a flow name, a location, and a unit. This metadata is useful if you want to edit or create new exchange links.
The Brightway2 exporter will automatically re-link (i.e. find the correct input values) exchanges when writing a new database. You can also manually create input values - no input value will be overwritten. In the database component of the input field, you can either use the name of the new database to be written, or the name of one of the input databases (it will be updated automatically).
Spatial relationships¶
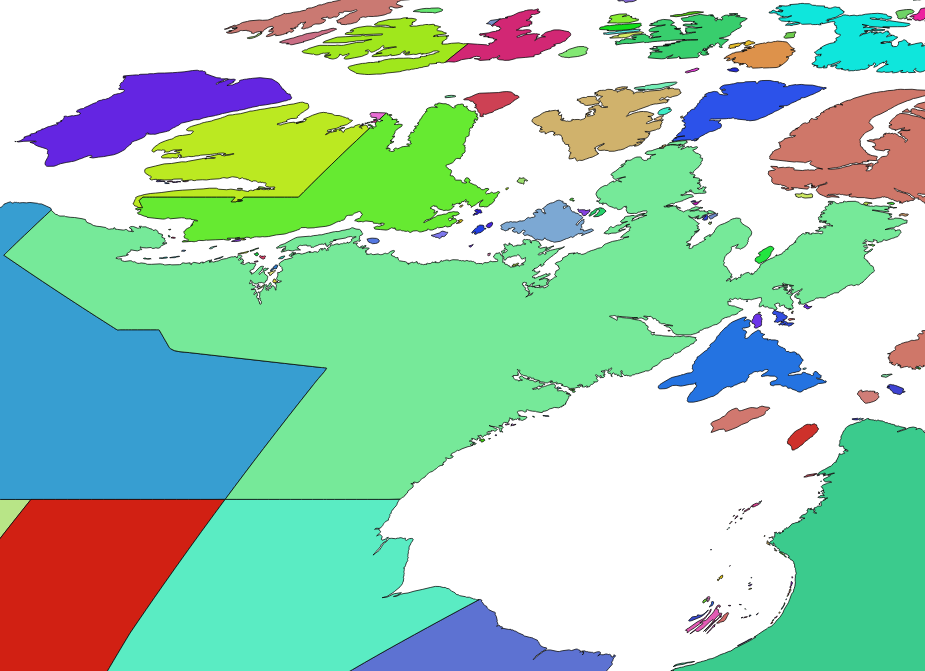
Topological faces in Northeastern Canada, showing both political and geographical divisions.
Wurst uses the constructive_geometries library to make spatial calculations easy. As shown above, constructive_geometries splits the world into a consistent set of topological faces, identified by integer ID values. This means that we can skip GIS functions like intersects, overlaps, etc. and instead use set algebra.
constructive_geometries is based on the natural earth database, and includes all countries, UN regions and subregions, some disputed areas, and a number of ecoinvent-specific regions; for full documentation, ; see the ecoinvent report for a complete list. Countries are identified by their two-letter ISO 3166-2 codes.
We recommend using the function relink_technosphere_exchanges, as this should be flexible enough for almost all cases, and has been tested to avoid common corner cases and possible hidden errors. See also the Matching and linking datasets in space example notebook.
-
wurst.transformations.geo.relink_technosphere_exchanges(ds, data, exclusive=True, drop_invalid=False, biggest_first=False, contained=True)¶ Find new technosphere providers based on the location of the dataset.
Designed to be used when the dataset’s location changes, or when new datasets are added.
Uses the name, reference product, and unit of the exchange to filter possible inputs. These must match exactly. Searches in the list of datasets
data.Will only search for providers contained within the location of
ds, unlesscontainedis set toFalse, all providers whose location intersects the location ofdswill be used.A
RoWprovider will be added if there is a single topological face in the location ofdswhich isn’t covered by the location of any providing activity.If no providers can be found, relink_technosphere_exchanes will try to add a RoW or GLO providers, in that order, if available. If there are still no valid providers, a
InvalidLinkexception is raised, unlessdrop_invalidisTrue, in which case the exchange will be deleted.Allocation between providers is done using
allocate_inputs; results seem strange ifcontained=False, as production volumes for large regions would be used as allocation factors.Input arguments:
ds: The dataset whose technosphere exchanges will be modified.data: The list of datasets to search for technosphere product providers.exclusive: Bool, default isTrue. Don’t allow overlapping locations in input providers.drop_invalid: Bool, default isFalse. Delete exchanges for which no valid provider is available.biggest_first: Bool, default isFalse. Determines search order when selecting provider locations. Only relevant isexclusiveisTrue.contained: Bool, default isTrue. If ture, only use providers whose location is completely within thedslocation; otherwise use all intersecting locations.
Modifies the dataset in place; returns the modified dataset.
Brightway2 IO¶
-
wurst.brightway.extract_database.extract_brightway2_databases(database_names, add_properties=False, add_identifiers=False)¶ Extract a Brightway2 SQLiteBackend database to the Wurst internal format.
database_namesis a list of database names. You should already be in the correct project.Returns a list of dataset documents.
-
wurst.brightway.write_database.write_brightway2_database(data, name)¶ Write a new database as a new Brightway2 database named
name.You should be in the correct project already.
This function will do the following:
- Change the database name for all activities and internal exchanges to
name. All activities will have the new databasename, even if the original data came from multiple databases. - Relink exchanges using the default fields:
('name', 'product', 'location', 'unit'). - Check that all internal links resolve to actual activities, If the
inputvalue is('name', 'bar'), there must be an activity with the codebar. - Check to make sure that all activity codes are unique
- Write the data to a new Brightway2 SQLite database
Will raise an assertion error is
namealready exists.Doesn’t return anything.
- Change the database name for all activities and internal exchanges to